|
Table of Contents
CellTrack: An Open-Source Software for Cell Tracking and Motility AnalysisMotivation: Cell motility is a critical part of many important biological processes. Automated and sensitive cell tracking is essential to cell motility studies where the tracking results can be used for diagnostic or curative decisions and where mathematical models can be developed to deepen the understanding mechanisms underlying the cell motility. Results: We propose a novel edge-based method for sensitive tracking of cells, and propose a scaffold of methods that achieves refined tracking results even under large displacements or deformations of the cells. The proposed methods along with other general purpose image enhancement methods are implemented in CellTrack, a self-contained, extensible, and cross-platform software package. Availability: CellTrack is an Open Source project and is freely available at http://sacan.biomed.drexel.edu/celltrack Contact: ahmet.sacan@drexel.edu Publication: If you use CellTrack for you own research, please cite the following paper:
Sample MoviesYou can download some sample movies and view the tracking results:
Download SoftwareDepending on your operating system, you need to download one of the following files:
Installation Instructions
On a linux-based system, you can download, uncompress, and compile CellTrack using the following commands: wget http://db.cse.ohio-state.edu/CellTrack/celltrack.tgz tar -xzf celltrack.tgz cd CellTrack # you may need to edit the Makefile to reflect the library paths make make install
There is also a Visual Studio project file provided under Please feel free to contact sacan@cse.ohio-state.edu if you have any compiling or installation questions. System Requirements
Quick Start GuideThe following are the common steps to be followed in CellTrack to perform a cell tracking:
LimitationsNote that CellTrack is by no means a final solution to all cell tracking problems. Depending on the movie you are processing, you might not get satisfactory results. We recommend that you go through each method and preview the effect of the method parameters on tracking results. The current version of CellTrack provides you with default parameters, but does not provide an automatic mechanism that can adjust the parameters to suit best the nature of the environment the cells are being tracked in or the conditions that your movie is captured. Furthormore, the current version of CellTrack does not handle occlusions. The occlusions are planned to be handled using modelling of the trajectory of individual snaxels using Kalman Filter and/or Condensation algorithm. Detailed User GuideHere we describe the available operations you can perform in CellTrack software. For some of the image processing operations, we have used the description of the parameters from the OpenCV documentation. The movie used for creating sample snapshots is copyright to the Cell Biology and Cytoskeleton Group at Harvard.
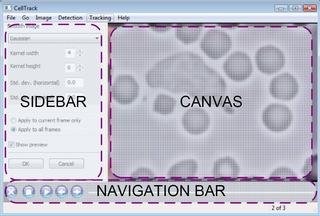
Below, you see a snapshot of the graphical user interface components. The current frame of the movie is shown in Common Plugin Sidebar Options
Menus and Shortcuts
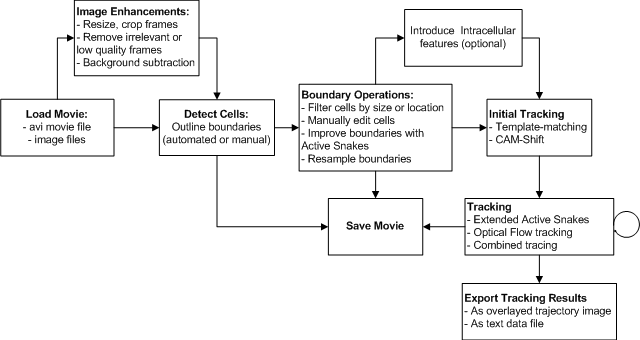
Task FlowchartLoading A Cell Movie
Commands: In CellTrack, your cell movies can either be an *.avi movie file or a set image files. CellTrack can read avi files using platform-specific video libraries. If you are having difficulty loading your movie file, please use a conversion tool to convert your movie into DIB/I420/IYUV format or uncompressed format. You can use one of the following freely available tools for this conversion: FFmpeg, mencoder, or RAD Video Tools. Instead of a movie file, you can also use a set of image files. Make sure the alphabetical order of the image file names are the same order of the real motion capture. The following image formats are supported:
Saving Movie or Frames
Commands:
You can save the whole movie (with object boundaries and intracellular components drawn) or individual frames. For saving individual frames, the following image formats are supported: bmp, png, jpeg, giv, pcx, pnm, xpm, ico, and cur. For saving the whole movie, you will only be able to save as an .avi movie file, but you can select the compression codec to be applied. The preferences dialog (File→Preferences Ctrl+K) has a list of available codecs you can choose from on the Navigation and Playback
Commands:
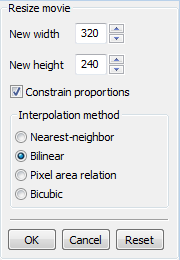
You can navigate through the loaded cell movie using the menu commands or using the navigation bar at the bottom of the window. When you are playing a movie file, the frame rate of the movie file is used for playback. When you are playing a set of image files, the default frame rate (5 f.p.s.) is used; the default frame rate can be changed in Basic Image OperationsIf you are working with large movies or images, we recommend that you reduce the size and frame count during the test phase. Once you have determined the best set of operations and parameters suited to your movie, you can apply them to the original data. Resize
Command:
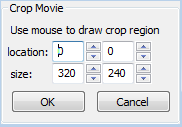
~~END_FLOAT_LEFT~~ Crop
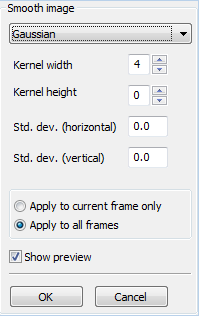
If you are concerned with only a limited subregion of the movie, you can crop the movie to use only this subregion. Cropping will remove the rest of the area and will save computational time for subsequent detection or tracking operations. You can use your mouse to draw the crop region on the canvas, or use the location and size input controls in the sidebar to define the region. Smooth
Command: You can perform smoothing to remove the noise in your images. The following smoothing methods are available:
~~END_FLOAT_LEFT~~ Removing frames
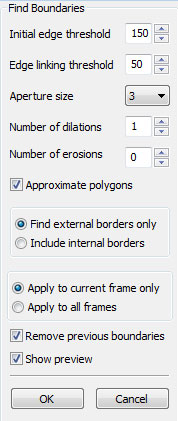
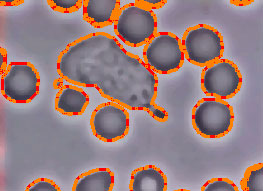
Commands: If you are only concerned with a specific section of the movie, you can remove the rest of the frames using the commands above. The tracking methods in CellTrack assume that the first frame includes the boundaries of the object you wish to track. If you define boundaries of interest in another frame, you need to delete the preceeding frames to force CellTrack to track these boundaries. You can also remove low-quality frames where the image is too noizy or blurry. If the movie capture is from a nonstable, moving camera, you might get some fuzzy transition frames which might need to be removed to obtain higher-quality tracking results. Automated Cell Detection
Filtering Cells
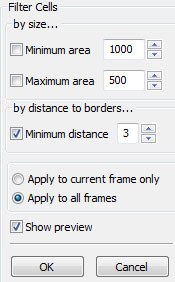
The automated cell detection may detect contours which may or may not be cells and may or may not be of interest to you. Fitering Cells is a quick way of removing unwanted objects. You can restrict the size of objects that are to be considered as cells, or specify a minimum distance to frame borders to filter out the cells that are not completely captured within the frame.
Manually Editing Cell Boundaries
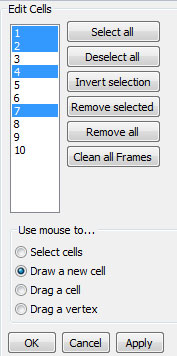
Use this plugin to manually outline a cell using the mouse. Click to start, move mouse to next vertex and click again. You can complete the drawing by either double-clicking at the next vertex, or clicking on the starting point. The sidebar will show the list of contours in the current frame. You can click on the cell id numbers to make selection. Use Shift, or Control keys in combination with mouse clicks to select multiple entries. You can then remove these contours from the current frame using
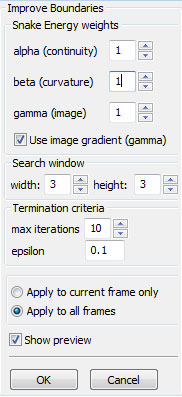
Choose from the Improving boundaries
Command:
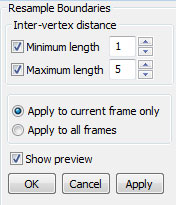
Resampling boundaries
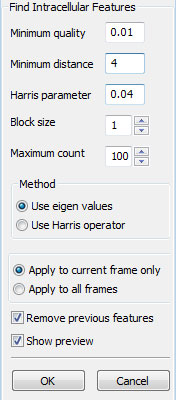
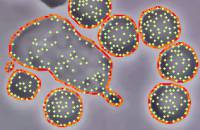
Introducing Intracellular Points
Command:
Tracking intracellular points would be useful if you are interested in the reorganization of the cell interior as it moves. CellTrack includes limited functionality for tracking intracellular points (features). The tracking methods that only shift/rotate the cells will perform tracking on the cell border and interior features equivalently and result in no conflicts. However, the optical-flow based tracking method is not interior-aware. Each of the border points and intracellular points are tracked independently, which may result in intracellular feeatures leaking out of the cell (The outlier detection and local interpolation may help avoid this conflict). The snake methods work only on the cell boundaries and do not modify or track intracellular points.
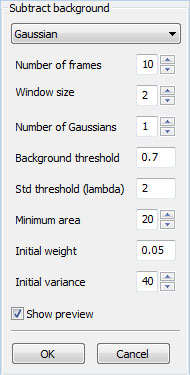
Note that, if you have high-resolution images where intracellular components (organelles or nucleus) are distinguishable, you can represent and track these components using internal boundaries instead of using intracellular points described in this plugin. Use Background Subtraction
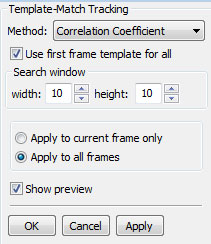
The Gaussian method is based on 'An Improved Adaptive Background Mixture Model for Real-time Tracking and Shadow Detection' by P. KaewTraKulPong and R. Bowden (Proceedings of the 2nd European Workshop on Advanced Video-Based Surveillance Systems, Sept. 2001). ~~END_FLOAT_RIGHT~~ Tracking methodsBelow are some of the common properties and parameters you need to be aware of when running the tracking plugins: Side-by-side display: You will see two canvas areas when you activate a tracking plugin. The canvas on the left will display previous frame, and the canvas on the right will display the next frame. If you are on the first frame currently, no frame will be shown on the left canvas; you will have navigate one frame forward. Search window: parameter limits the neighborhood that is searched in the new frame (around the location of the bounding rectangle in the previous frame). You need to use bigger search windows if the cells move at a very high speed (or if there is a shift in camera location – or equivalently the media is moved). A very low frame-rate would also result in having a larger shift in the location of the cells, requiring a bigger search window. Use avilable tracking, if any: This option, if selected, will initialize the tracking method with any boundaries or feature points already available in the current frame (The number of points must be the same as that of the previous frame, otherwise this option is ignored). Otherwise, an exact copy of the previous features is used as the initial guess. Template-matching based tracking
Command:
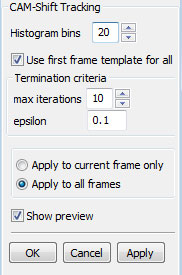
CAM-Shift tracking
Continuously Adaptive Mean SHIFT (CAM-Shift) tracking algorithm is an implementation of 'Real Time Face and Object Tracking as a Component of a Perceptual User Interface' by Gary R. Bradski (1998, Fourth IEEE Workshop on Applications of Computer Vision (WACV'98) p. 214). CAM-Shift is based on the mean shift algorithm ('Mean shift: a robust approach toward feature space analysis' by Comaniciu and Meer, IEEE Trans. on Pattern Analysis and Machine Intelligence, 1997), a robust non-parametric iterative technique for finding the mode of probability distributions.
CAM-Shift first finds the center of the cell in the new frame using the back projection of the histogram generated from the previous frame. Then it calculates the new size and orientation for the cell. Optical Flow tracking
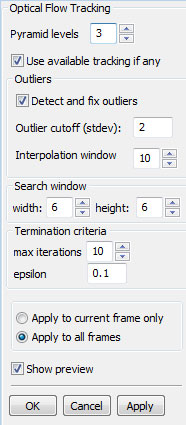
Command:
Note that the optical flow method itself does not guarantee that the relative locations of the points will be preserved during tracking. This may result in inconsistent tracking results along the boundaries, or leakage of intra-cellular features to outside the cell. Moreover, the optical flow method may even loose some of the features (i.e., these features are not found in the new frame). To remedy these problems, we have implemented an outlier detection and update mechanism. The points that move a distance that is above a certain threshold, which is specified as a user defined coefficient of the standard deviation of all the shifts ( Extended Snake tracking
Extended Snake tracking implements a new energy functional we have developed for sensitive tracking of cells. The additional energy terms help to match up the new snake with the old snake (that is, the one from the previous frame) both internally (in relation to itself) and externally (in relation to the image plane). See the section on Improving Boundaries for the explanation of the original snake parameters (alpha, beta, and gamma) used for improving a boundary; these parameters are used without any changes here. The
To illustrate the
Note: The Extended Snake Tracking method is currently being validated. An extensive quantitative evaluation along with comparison with other methods are currently in process. Combined tracking
Command:
You can select which methods to apply. An initial tracking can be calculated using either Template-matching or CAM-Shift methods. The Template-matching or CAM-Shift methods provide an initial tracking that is robust to large shifts of media or camera (We recommend you turn off Template-matching and CAM-Shift methods if your movie contains highly similar cells that are very close to each other). This initial method can then be improved by Optical Flow and/or Extended Snake tracking. The parameters for each of these methods can be adjusted within this plugin. After tracking at each new frame, you can select to
Note: The Combined Tracking method is currently being validated. An extensive quantitative evaluation along with comparison with other methods are currently in process.
Analyzing Tracking ResultsCellTrack provides some basic motility analysis features like ability to view the trajectory, speed, area, and deformation of the cells across movie frames. To make analysis using external programs possible, you can also export the tracking data as clear-text files. The tracking data of a cell is included only if this cell is present in the first frame of the movie. If you have frames at the beginning of the movie, you can remove these frames and make the starting frame of interest to be the first frame. Cell Speed
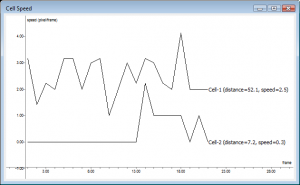
The speed of a cell is calculated as the displacement of its center of mass across frames. The figure on the right gives the speed plot of two cells. Cell 1 is a constantly moving cell, whereas Cell 2 starts moving at 10th frame and comes back to rest. The total distance traveled by each cell and the average speeds are also given in the plot legends. You can use the mouse to navigate in a plot. Right-click on the plot once to see available mouse commands. Right-click and drag the mouse to shift the plot.
If you wish to analyze the speed data for externally, or save it for future reference, you can use the #width: 320, height: 240, frameCount: 22, fps: 2 #cellCount: 2 #Cell: 1, pointCount: 168, totalDistance: 50.731670, avgSped: 2.415794 2.000000 2.828427 2.000000 ... #Cell: 2, pointCount: 60, totalDistance: 7.236068, avgSped: 0.344575 0.000000 0.000000 0.000000 ... ~~END_FLOAT_RIGHT~~ Area of the Cells
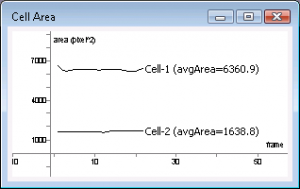
You can plot or export the area, or the change in the area, of the cells across frames. You can use the area information to filter cells based on their area. The area of the moving cells would change slightly as they move, whereas the area of the stable cells will not change. Note that significant changes in the area of a cell might be an indication of the cell moving out of focal plane or out of the view. The area plots and the data export format are similar to those of the Cell Speed described above. The area values are given in units of pixel^2. #width: 320, height: 240, frameCount: 22, fps: 2 #cellCount: 2 #Cell: 1, pointCount: 168, avgArea: 6342.636230 6666.000000 6478.500000 6362.500000 ... #Cell: 2, pointCount: 60, avgArea: 1622.250000 1618.000000 1639.500000 1580.500000 ... ~~END_FLOAT_LEFT~~ Cell Deformation
The deformation is calculated as the difference in the overlapped areas of the cells in consecutive frames. The cell from frame #width: 320, height: 240, frameCount: 22, fps: 2 #cellCount: 2 #Cell: 1, pointCount: 168, avgDeformation: 2012.023804 2031.500000 1988.000000 2000.000000 ... #Cell: 2, pointCount: 60, avgDeformation: 724.166687 713.500000 737.000000 727.500000 ... ~~END_FLOAT_RIGHT~~ Cell Trajectory
Commands: The trajectory gives only the tracking of the center of each cell, providing information about the overall movement of the cell. You can view or save the trajectory as an image, or export the trajectory data into a text file. For reference, the starting and ending configurations of the cell are also shown on the trajectory image. The speed of the whole cell described above is calculated using the amount of displacement between frames. The data format for the trajectory is similar to that of cell_speed. For each cell, tab-separated <x,y> coordinates of the center of mass is given on a separate line for each frame. The data sample below shows that the cell-1 (its center of mass) moves from <132,87> to <130,87> to <128,85>,…. #width: 320, height: 240, frameCount: 22, fps: 2 #cellCount: 2 #Cell: 1, pointCount: 168 132 87 130 87 128 85 ... #Cell: 2, pointCount: 60 164 30 164 30 164 30 ... ~~END_FLOAT_LEFT~~ Detailed Cell Tracking
Commands: The tracking image shows the path followed by each tracked point across frames. The beginning and the final configuration of the cell boundary are also drawn for reference. You can also export the tracking information as a data text file. The following shows a sample exported tracking data. For each cell, the object identifier and the number of control points on this cell are printed, followed by the x and y coordinates of each control point across all frames. For instance, the first control point of the first cell below moves from <119,41> to <118,42> to <118,41> in the first 3 frames. The coordinates are separated by tabs ('\t'). Note that the exported tracking data can later be imported into CellTrack. #width: 320, height: 240, frameCount: 22, fps: 2 #cellCount: 2 #Cell: 1, pointCount: 168 119 41 116 42 114 43 ... 118 42 116 43 115 44 ... 118 41 115 42 114 43 ... ... #Cell: 2, pointCount: 60 155 8 153 9 151 10 ... 155 8 153 9 151 10 ... 155 9 153 10 150 11 ... The following sample C++ code is provided for your convenience if you wish to parse the exported tracking data: FILE *fp = fopen("track.txt","r");
int NumCells;
int Width, Height, FrameCount, FSP;
if( fscanf(fp, "#width: %d, height: %d, frameCount: %d, fps: %d\n#cellCount: %d\n", &Width, &Height, &FrameCount, &FSP, &NumCells) !=5 ) {
wxLogError("Unable to parse file header %s", file); return false;
}
for (int c=0; c<NumCells; c++){
int cellid,np;
if( fscanf(fp, "#Cell: %d, pointCount: %d\n", &cellid, &np) !=2 ) {
wxLogError("Unable to parse file %s for cell %d header %s", file, c+1); return false;
}
for (int j=0; j<FrameCount; j++) {
std::vector<wxPoint> roi(np);
for (int i=0; i<np; i++){
if( fscanf(fp, "%d %d", &roi[i].x, &roi[i].y) !=2 ) {
wxLogError("Unable to parse file %s for cell %d coordinates", file, c+1); return false;
}
}
// the roi variable here contains the contour for Cell 'c'
// at frame 'j'
book[j]->AddRoi(roi);
}
}
fclose(fp);
~~END_FLOAT_RIGHT~~
Notes for Developers
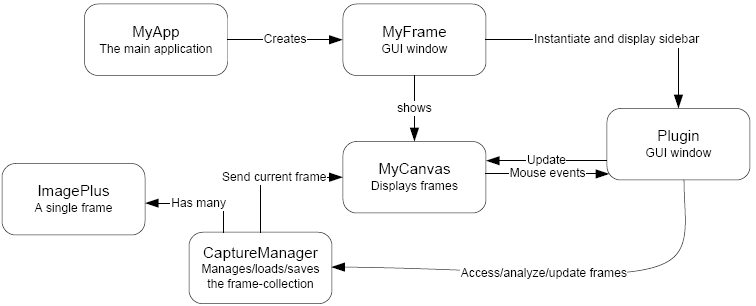
This section is geared toward researchers or developers wishing to extend CellTrack by implementing new plugins. A general overview of the source architecture is given followed by instructions to get you started in writing a new plugin. The figure below shows a simplified outline of the classes and modules used in CellTrack.
If you want to develop a new plugin, you will need to first design a sidebar to accept the parameters (if any) from the user. See
The plugin module itself needs to be a subclass of
The previewing and committing of changes to all frames is automatically performed by the base class. You may want to change the default processing by overriding the relavant functions (such as
In most cases, you won't need to access the Canvas directly. However, if you are implementing a plugin where you need to gather information through user's actions on the canvas (such as dragging a cell using the mouse), you need to register the plugin as a listener of the relavant events. Please see the |